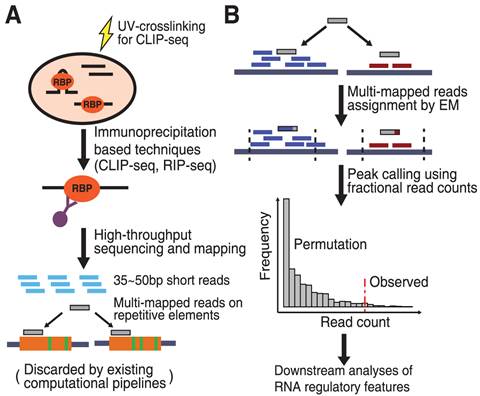
Mapping transcriptome-wide protein-RNA interactions to elucidate RNA regulatory programs. - Abstract - Europe PMC

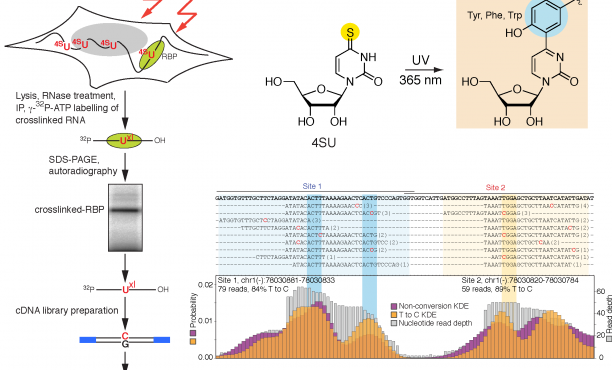
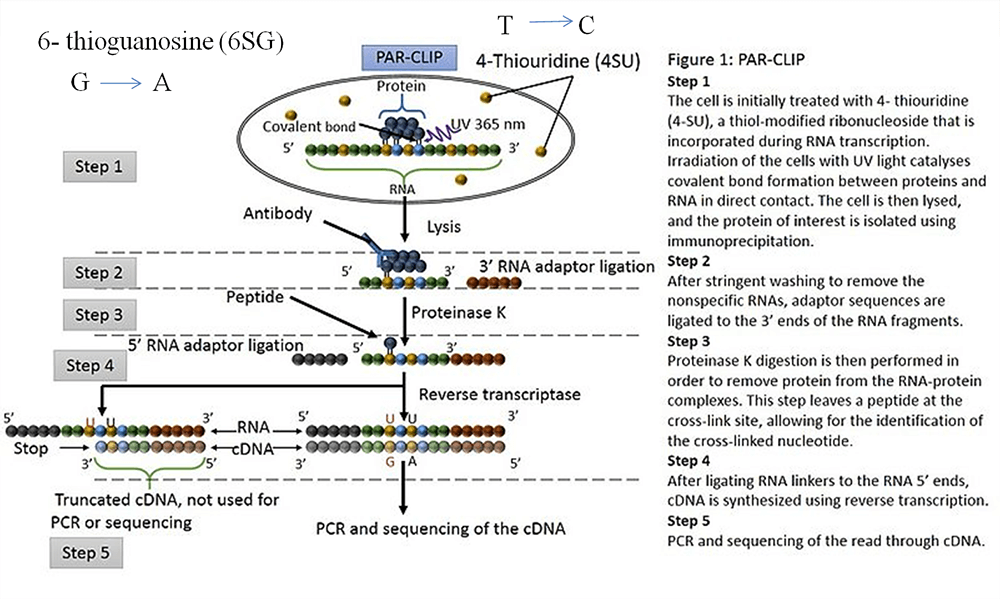
Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

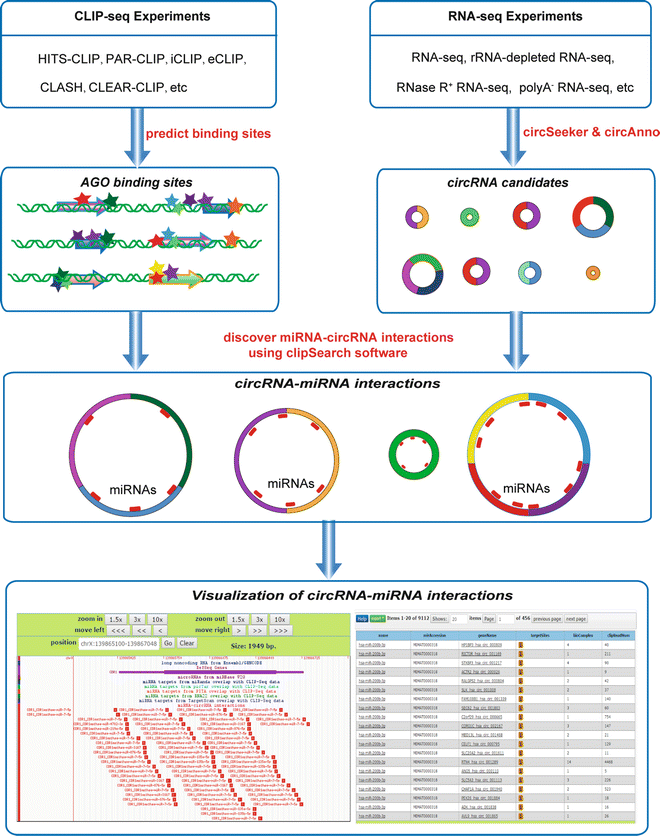
starBase v2.0: decoding Interaction Networks of lncRNAs, miRNAs, ceRNAs, RNA-Binding Proteins and mRNAs from large-scale tumor samples and CLIP-Seq (HITS-CLIP, PAR-CLIP, iCLIP, CLASH) data
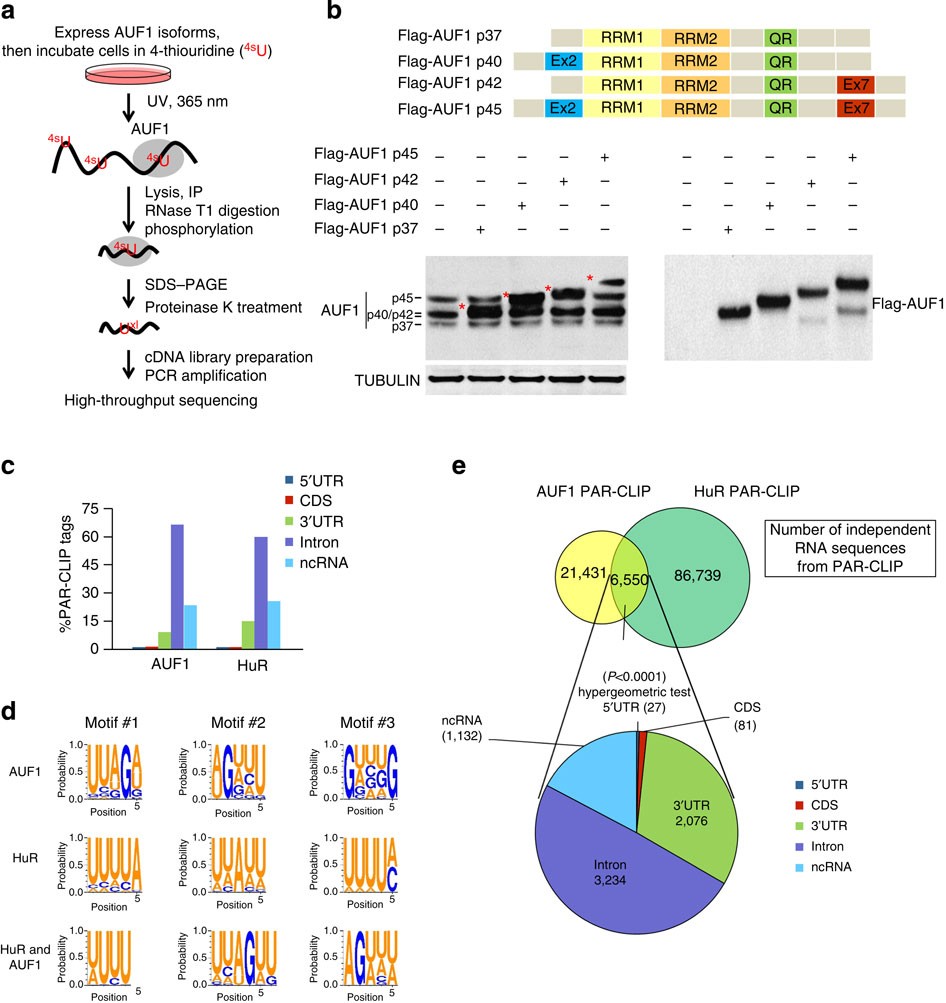
PAR-CLIP analysis uncovers AUF1 impact on target RNA fate and genome integrity | Nature Communications
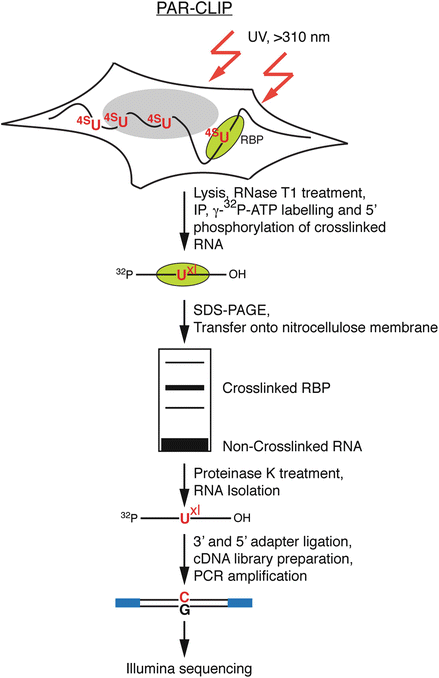
PAR-CLIP: A Method for Transcriptome-Wide Identification of RNA Binding Protein Interaction Sites | SpringerLink

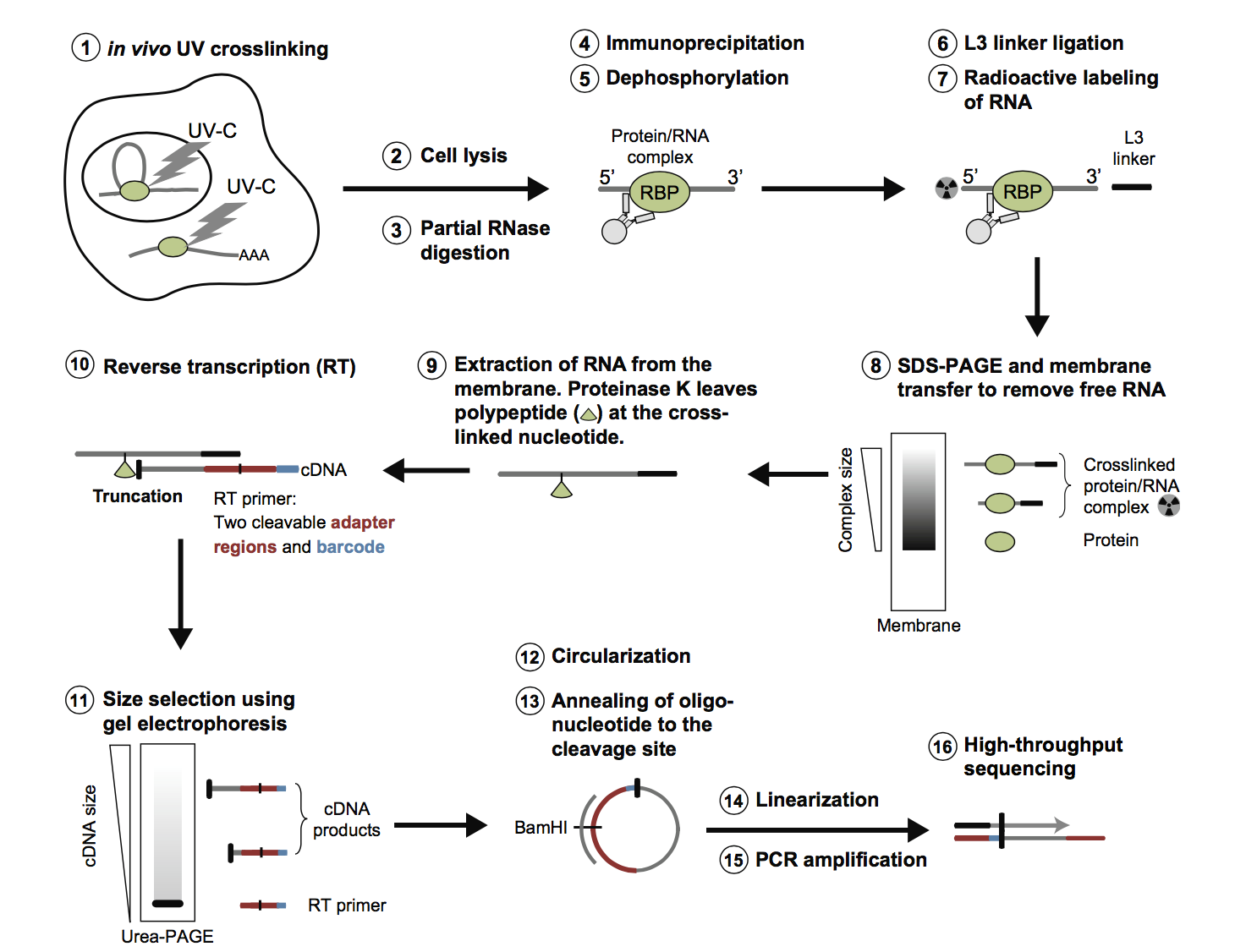
PAR-CLIP and streamlined small RNA cDNA library preparation protocol for the identification of RNA binding protein target sites - ScienceDirect
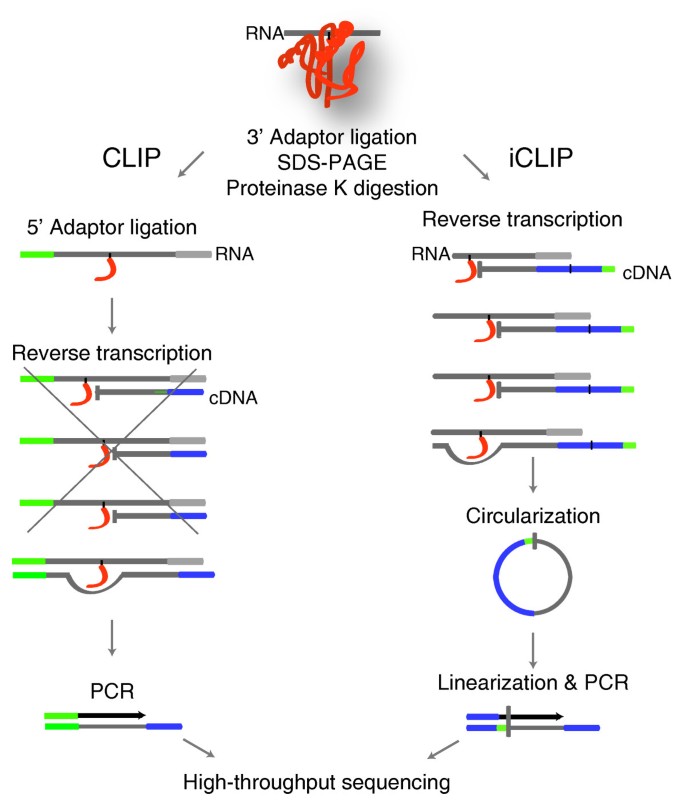
Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text
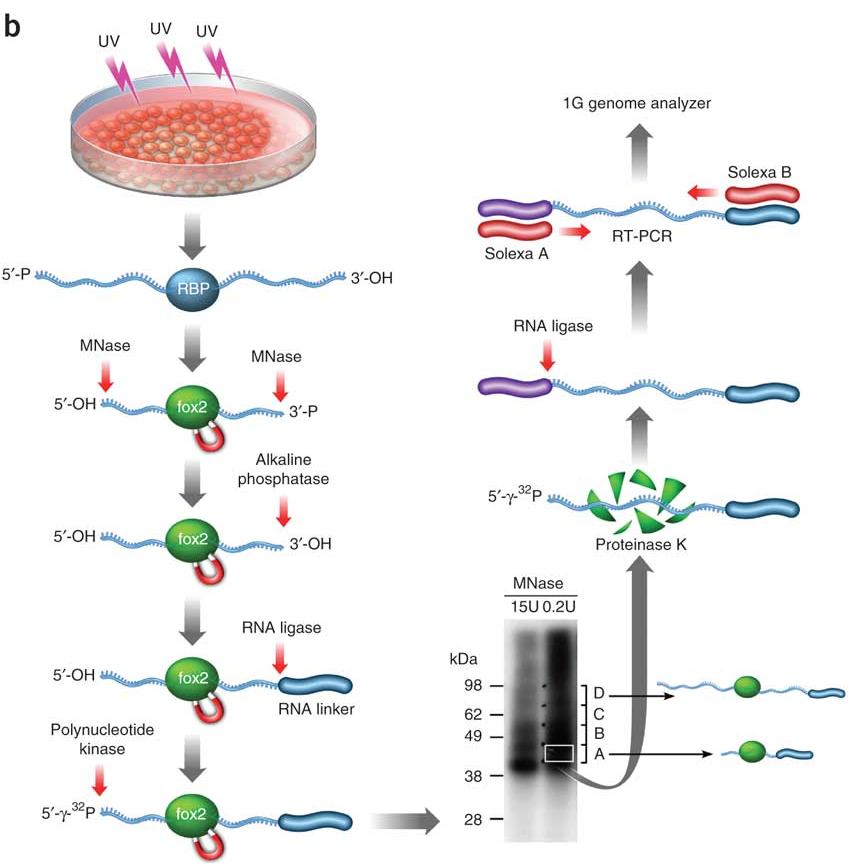
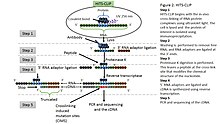
Identify RNA-RBP Direct Interactions in Genome-wide Scale Fast Introduction to HITS-CLIP (or CLIP-seq) Yun YAN Press Spacebar or Tab To Get Started HITS-CLIP (or CLIP-seq) HIThroughput Sequencing of RNAs isolated by CrossLinking ...

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

Optimization of PAR-CLIP for transcriptome-wide identification of binding sites of RNA-binding proteins - ScienceDirect

PAR-CliP - A Method to Identify Transcriptome-wide the Binding Sites of RNA Binding Proteins | Protocol

Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram